| |
 |
Protein Localization Database for Human and Arabidopsis |
|
|
| |
|
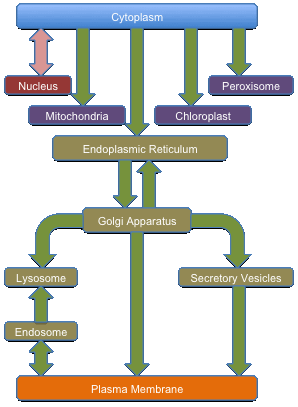
Protein Subcellular LocalizationLocDB is a expert curated database that collects experimental annotations for the subcellular localization of proteins in Human (Homo sapiens) and Weed (Arabidopsis thaliana). The database also contains predictions of subcellular localization from a variety of state-of-the-art prediction methods for all proteins with experimental information. Proteins are the fundamental functional components of cells. They are responsible for transforming genetic information into physical reality. These macromolecules mediate gene regulation, enzymatic catalysis, cellular metabolism, DNA replication, and transport of nutrients, recognition, and transmission of signals. The interpretation of this wealth of data to elucidate protein function in post-genomic era is the a fundamental challenge. To date, even for the most well-studied organisms such as yeast, about one-fourth of the proteins remain uncharacterized. Major obstacle in experimentally determining protein function is that the studies require enormous resources. Hence, the gap between the amount of sequences deposited in databases and the experimental characterization of the corresponding proteins is ever-growing. Bioinformatics plays a central role in bridging this sequence-function gap through the development of tools for faster and more effective prediction of protein function. With this repository we are trying to fill the gap between experimental annotations and predictions and providing a bigger dataset for the testing of new prediction methods.
Who are we?Rostlab: The Rost Group at TU Munich |
|